Problem Set 4 KEY
8) Estimate Vmax and Km for the following data
|
[S] mM |
v mM-min-1 |
|
5 |
22 |
|
10 |
39 |
|
20 |
65 |
|
50 |
102 |
|
100 |
120 |
|
200 |
135 |
using:
a. Plot of v versus [S]
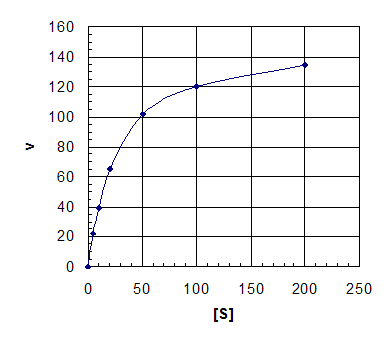
Eyeballing, can guess curve flattens out at
about Vmax = 150 mM/min, in which case Km is about 25 mM.
a. Lineweaver-Burke

Vmax = 1/0.0061 = 164;
Km/Vmax = 0.1961, so Km = 164*.1961 = 32
b. Eadie-Hofstee
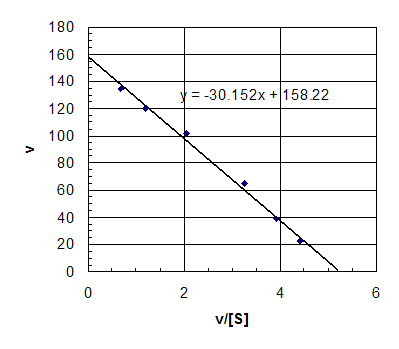
Km is about 30, Vmax
is about 158
9) Total enzyme concentration in (8) is 1 nM.
a. How many substrate molecules can an enzyme catalyze in 1 minute?
If we average the results from b and c above,
we get Vmax = 161mM/min and Km = 31 mM. Assuming those
values to be accurate, then:
![]()
b. Calculate kcat/Km for the enzyme
kcat is usually
expressed in terms of sec-1, not min-1, co we need to convert (a):
![]()
kcat/Km is then:
![]()
This is reasonably efficient, though not at
the top end of enzyme efficiencies (>108).
15) Kinetics of an enzyme are studied with and without an inhibitor, giving the following results:
|
S (mM) |
V- (mM/min) |
V+ (mM/min) |
1/S |
1/V- |
1/V+ |
|
1.25 |
1.72 |
0.98 |
0.8 |
0.581395 |
1.020408 |
|
1.67 |
2.04 |
1.17 |
0.598802 |
0.490196 |
0.854701 |
|
2.5 |
2.63 |
1.47 |
0.4 |
0.380228 |
0.680272 |
|
5 |
3.33 |
1.96 |
0.2 |
0.3003 |
0.510204 |
|
10 |
4.17 |
2.38 |
0.1 |
0.239808 |
0.420168 |
a. What kind of inhibitor?
b. kcat and Km with and without inhibitor
Okay – let’s plot the data:
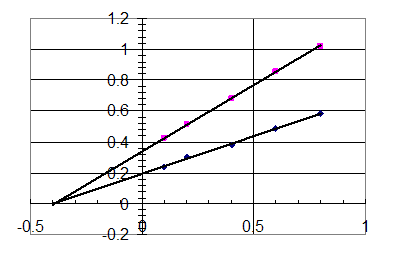
Since Vmax changes
and Km is constant, this is noncompetitive inhibition.
Km = 2.5 mM
Vmax without inhibitor
is about 5 mM/min
Vmax with inhibitor is
about 3 mM/min
16) The same enzyme as in Problem 15 is analyzed in the presence of another inhibitor at two inhibitor concentrations (3 mM and 5 mM) with the following results. (Shaded numbers were calculated from the unshaded numbers as part of the solution.)
|
S |
V - I |
V + 3mM I |
V + 5mM I |
1/S |
1/V- |
1/V+3mM |
1/V+5mM |
|
1.25 |
1.72 |
1.25 |
1.01 |
0.80 |
0.58 |
0.80 |
0.99 |
|
1.67 |
2.04 |
1.54 |
1.26 |
0.60 |
0.49 |
0.65 |
0.79 |
|
2.5 |
2.63 |
2 |
1.72 |
0.40 |
0.38 |
0.50 |
0.58 |
|
5 |
3.33 |
2.86 |
2.56 |
0.20 |
0.30 |
0.35 |
0.39 |
|
10 |
4.17 |
3.7 |
3.49 |
0.10 |
0.24 |
0.27 |
0.29 |
We can display the results on a Lineweaver-Burke plot (an Eadie-Hofstee plot would work just as well).
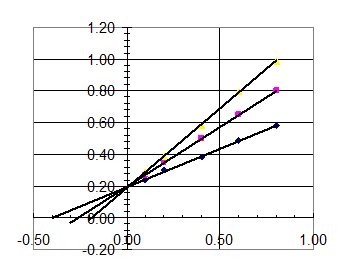
Regression of the resulting lines gives the
following:
|
0mM inhib |
y =
0.4843x + 0.1951 |
|
|
3mM inhib |
y =
0.755x + 0.1969 |
|
|
5 mM inhib |
y =
1.0062x + 0.1861 |
|
a)
Vmax is unchanged; Km varies with inhibitor
concentration. Therefore this is competitive inhibition.
b)
Kmapp’s (apparent Km in
the presence of inhibitor) and Vmax are calculated
from the plot as:
|
Vmax |
5. |
|
Km 0 mM
I |
2.5 |
|
Km 3 mM
I |
3.8 |
|
Km 5 mM
I |
5.4 |
c)
KI can be determined from the apparent Km’s and the
relationship between Km and Kmapp for
competitive inhibitors (see eq. 11.37a and fig.
11.20c).
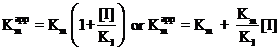
We
plot Kmapp vs. [I]. The y-intercept is Km; the slope id Km/KI.
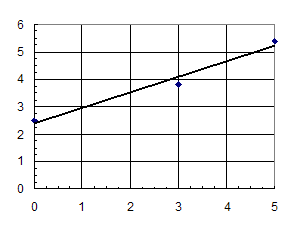
|
Km |
2.5 |
|
Km/Ki |
0.5684 |
|
Ki |
4.4 |
Numbers may vary slightly depending on your
plot.
20) The following data are obtained for the steady state kinetics of an enzyme-catalyzed reaction (shaded values are calculated):
|
[S] mM |
v mM/sec |
1/[S] |
1/v |
|
0.250 |
0.260 |
4.000 |
3.846 |
|
0.330 |
0.450 |
3.030 |
2.222 |
|
0.500 |
0.920 |
2.000 |
1.087 |
|
0.750 |
1.800 |
1.333 |
0.556 |
|
1.000 |
2.500 |
1.000 |
0.400 |
|
2.000 |
4.100 |
0.500 |
0.244 |
|
4.000 |
4.800 |
0.250 |
0.208 |
a) Does the enzyme exhibit Michaelis-Menten kinetics?
We can make a Lineweaver-Burke
plot; the result is not linear, so Michaelis-Menten
kinetics are violated.
(It is left as an exercise for the student to plot v versus [S] – the
resulting plot is slightly sigmoidal, implyimg cooperativity.)
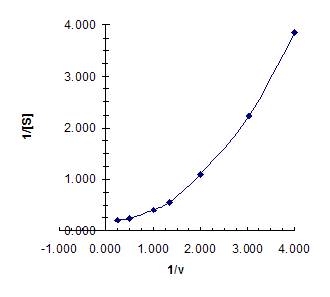
b) What is Vmax?
My best eyeball guess from the above plot is
that – extrapolating the curve – we get a y-intercept (corresponding to 1/Vmax)
of about 0.2 sec/mM, giving Vmax of about 5 mM/sec.